Autoencoder on MNIST¶
Example for training a centered Autoencoder on the MNIST handwritten digit dataset with and without contractive penalty, dropout, …
It allows to reproduce the results from the publication How to Center Deep Boltzmann Machines. Melchior et al. JMLR 2016..
Theory¶
If you are new on Autoencoders visit Autoencoder tutorial or watch the video course by Andrew Ng.
Results¶
The code given below produces the following output that is quite similar to the results produced by an RBM.
Visualization of 100 test samples.
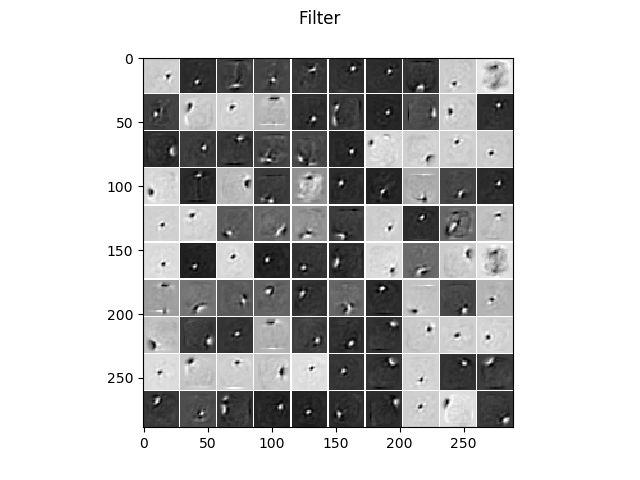
The learned filters without regularization.
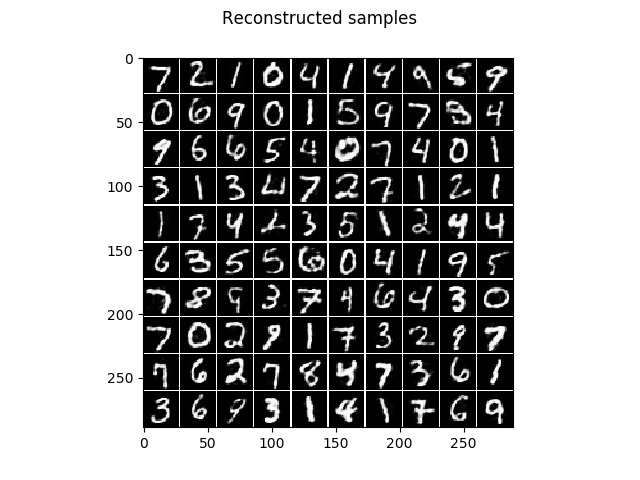
The corresponding reconstruction of the model, that is the encoding followed by the decoding.
The learned filters when a contractive penalty is used, leading to much more localized and less noisy filters.
And the corresponding reconstruction of the model.
See also RBM_MNIST_big.
Source code¶
""" Example for contractive Autoencoder (SAE) on MNIST.
:Version:
1.0.0
:Date:
28.01.2018
:Author:
Jan Melchior
:Contact:
JanMelchior@gmx.de
:License:
Copyright (C) 2018 Jan Melchior
This file is part of the Python library PyDeep.
PyDeep is free software: you can redistribute it and/or modify
it under the terms of the GNU General Public License as published by
the Free Software Foundation, either version 3 of the License, or
(at your option) any later version.
This program is distributed in the hope that it will be useful,
but WITHOUT ANY WARRANTY; without even the implied warranty of
MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the
GNU General Public License for more details.
You should have received a copy of the GNU General Public License
along with this program. If not, see <http://www.gnu.org/licenses/>.
"""
# Import numpy, i/o functions, preprocessing, and visualization.
import numpy as numx
import pydeep.misc.io as io
import pydeep.misc.visualization as vis
import pydeep.preprocessing as pre
# Import cost functions, activation function, Autencoder and trainer module
import pydeep.base.activationfunction as act
import pydeep.base.costfunction as cost
import pydeep.ae.model as aeModel
import pydeep.ae.trainer as aeTrainer
# Set random seed (optional)
numx.random.seed(42)
# Input and hidden dimensionality
v1 = v2 = 28
h1 = 10
h2 = 10
# Load data , get it from 'deeplearning.net/data/mnist/mnist.pkl.gz'
train_data, _, _, _, test_data, _ = io.load_mnist("mnist.pkl.gz", False)
# Set hyperparameters batchsize and number of epochs
batch_size = 10
max_epochs = 10
# Create model with sigmoid hidden units, linear output units, and squared error loss.
ae = aeModel.AutoEncoder(v1*v2,
h1*h2,
data = train_data,
visible_activation_function = act.Sigmoid(),
hidden_activation_function = act.Sigmoid(),
cost_function = cost.CrossEntropyError(),
initial_weights = 'AUTO',
initial_visible_bias = 'AUTO',
initial_hidden_bias = 'AUTO',
initial_visible_offsets = 'AUTO',
initial_hidden_offsets = 'AUTO',
dtype = numx.float64)
# Initialized gradient descent trainer
trainer = aeTrainer.GDTrainer(ae)
# Train model
print 'Training'
print 'Epoch\tRE train\t\tRE test\t\t\tSparsness train\t\tSparsness test '
for epoch in range(0,max_epochs+1,1) :
# Shuffle data
train_data = numx.random.permutation(train_data)
# Print reconstruction errors and sparseness for Training and test data
print epoch, ' \t\t', numx.mean(ae.reconstruction_error(train_data)), ' \t',\
numx.mean(ae.reconstruction_error(test_data)), ' \t', \
numx.mean(ae.encode(train_data)), ' \t',\
numx.mean(ae.encode(test_data))
for b in range(0,train_data.shape[0],batch_size):
trainer.train(data = train_data[b:(b+batch_size),:],
num_epochs=1,
epsilon=0.1,
momentum=0.0,
update_visible_offsets=0.0,
update_hidden_offsets=0.01,
reg_L1Norm=0.0,
reg_L2Norm=0.0,
corruptor=None,
reg_sparseness = 0.0,
desired_sparseness=0.0,
# Set to 0.0 to disable contractive penalty
reg_contractive=0.3,
reg_slowness=0.0,
data_next=None,
restrict_gradient=0.0,
restriction_norm='Cols')
# Show filters/features
filters = vis.tile_matrix_rows(ae.w, v1,v2,h1,h2, border_size = 1,
normalized = True)
vis.imshow_matrix(filters, 'Filter')
# Show samples
samples = vis.tile_matrix_rows(test_data[0:100].T, v1,v2,10,10,
border_size = 1,
normalized = True)
vis.imshow_matrix(samples, 'Data samples')
# Show reconstruction
samples = vis.tile_matrix_rows(ae.decode(ae.encode(test_data[0:100])).T,
v1,v2,10,10,
border_size = 1,
normalized = True)
vis.imshow_matrix(samples, 'Reconstructed samples')
# Show all windows.
vis.show()